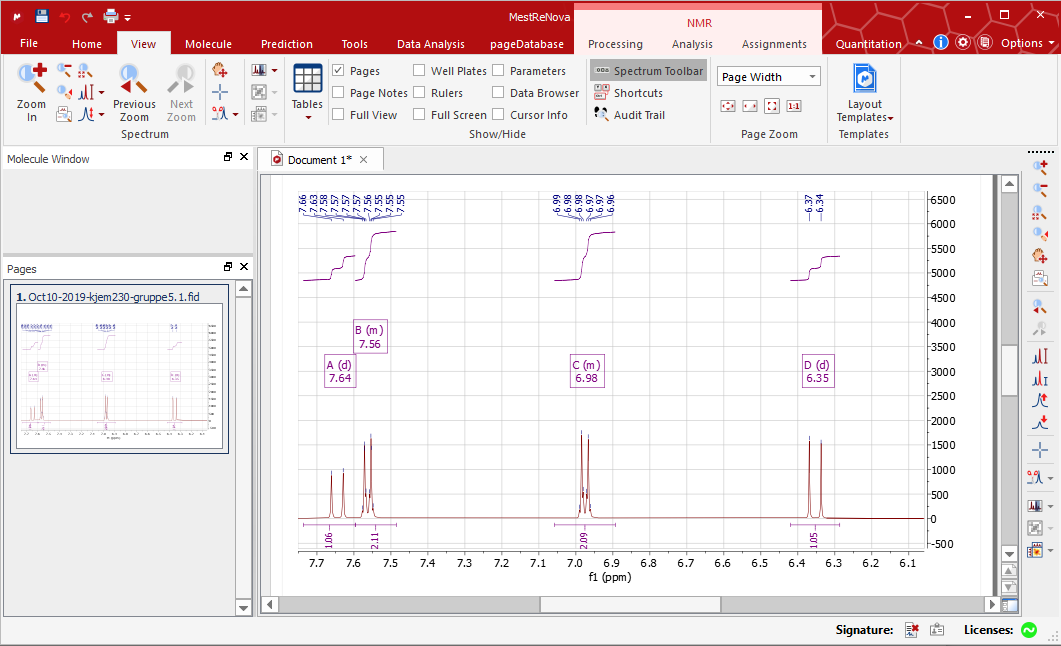
Unfortunately the multiplet integral is calculated automatically, and while.
#MESTRENOVA LABEL LICENSE#
Again select your license and push the button labeled Activate Selected. The multiplet integral does show up when you stack spectra. All new NMR users at Notre Dame should start with MNova Lite or Topspin. Then from the text toolbar (by default is usually at the bottom of the screen, but if not shown go to >View>Toolbars>Text) change the font size to something small enough to fit on the screen. Depending on what version you have (I think it needs to be MNova 9 or later), you can use the multiplet function (default shortcut J) and it will display an integral for the region (by default a purple number). Make sure under the geometry tab, the Auto box is selected for size.
#MESTRENOVA LABEL MANUAL#
You may wish to play with the manual multiplet buttons, to hover over the integral lines, right-click, and Edit the Multiplet to change its description, the number of protons you believe it corresponds to, or the peaks selected. Once Mnova has detected multiplets, you should look at them. Left click the parameters (which should have the green selected boxes shown) and choose properties. that Mnova labels the water and residual DMSO-d 5 signals. You need to resize the text to a smaller font size Double click on the spectrum display and select Properties/Multiplets to display the Multiplets Properties dialog box Once there, you can choose the color, the font and the margin of the Multiplet boxes. This title appears by default on each page of the page navigator. Bear in mind that, Mnova will only show the first line of the original title. Left click and hold to define a box within which you will have your peaks pickedģ)how on earth do you get the parameter box to fit onto the page? Ive managed to get it up but it is about three times larger than my spectra and rescaling using the little green buttons leaves it the same size but just cuts loads out. Multiplet Label By Administrator on 6 May, 2010 Resource It is really easy to customize the multiplet boxes. The user can include the spectrum title (the title set previously on the spectrometer at acquire time) on any axis label by typing 't' in its corresponding edit box. >Analysis >Peak Picking >Manual Threshold (or just type K) Mnova can analyze and report multiplets in terms of chemical shift, splitting pattern, J-coupling(s), and of protons. >View >Zoom >Manual Zoom (or just type M, or click the manual zoom button)Ģ)Set a threshold so that only peaks above a certain intensity will automatically label. Testing this on version 9 will have to wait until tomorrow, but on version 8 (which I have in front of me, but which has limited differences to version 9 by memory)ġ)Rescale the chemical shift axis to focus only on the relevant sections of my spectra.
